R Tools for Research Publication
Reproducibe Research
When writing a manuscript or developing a data story, I use RMarkdown. RMarkdown enables authors to prepare reproducible research documents, which
ties specific instructions to data analysis and experimental data so that scholarship can be recreated, better understood and verified. Packages in R for this purpose can be split into groups for: literate programming, package reproducibility, code/data formatting tools, format convertors, and object caching. The primary way that R facilitates reproducible research is using a document that is a combination of content and data analysis code. The Sweave function (in the base R utils package) and the knitr package can be used to blend the subject matter and R code so that a single document defines the content and the analysis.
A great graphic that illustrates the use of various R packages to develop a RMarkdown document is found at the Aberdeen Study Group.
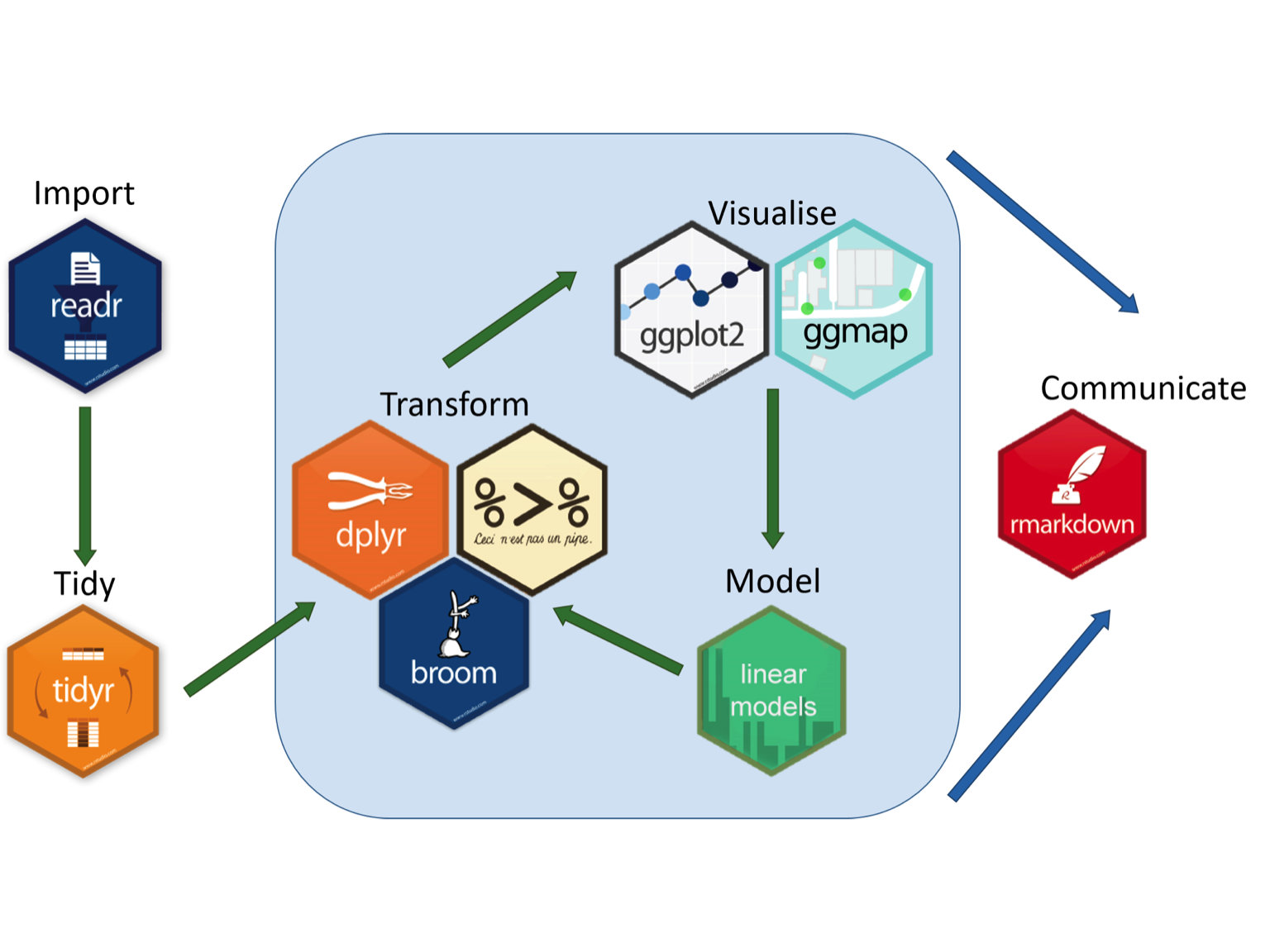
For blogging, I use Yihui Xie’s blogdown approach, with Hugo themes, hosted on private Github repository that is deployed to Netlify. It may seem a little complex, but I can assure you it isn’t. I plan, however, to prepare an in depth video on how I do this at a later time.
Here, I share the finer details to streamline the manuscript preparation or data story process.
RMarkdown
RMarkdown, created in RStudio, is a productive notebook interface to weave together narrative text and code to produce elegantly formatted output. Multiple coding languages including R, Python, and SQL can be used in an RMarkdown document. When you get a chance, checkout the RMarkdown gallery. In RStudio, you can create a new RMarkdown by clicking the new file icon as shown here:
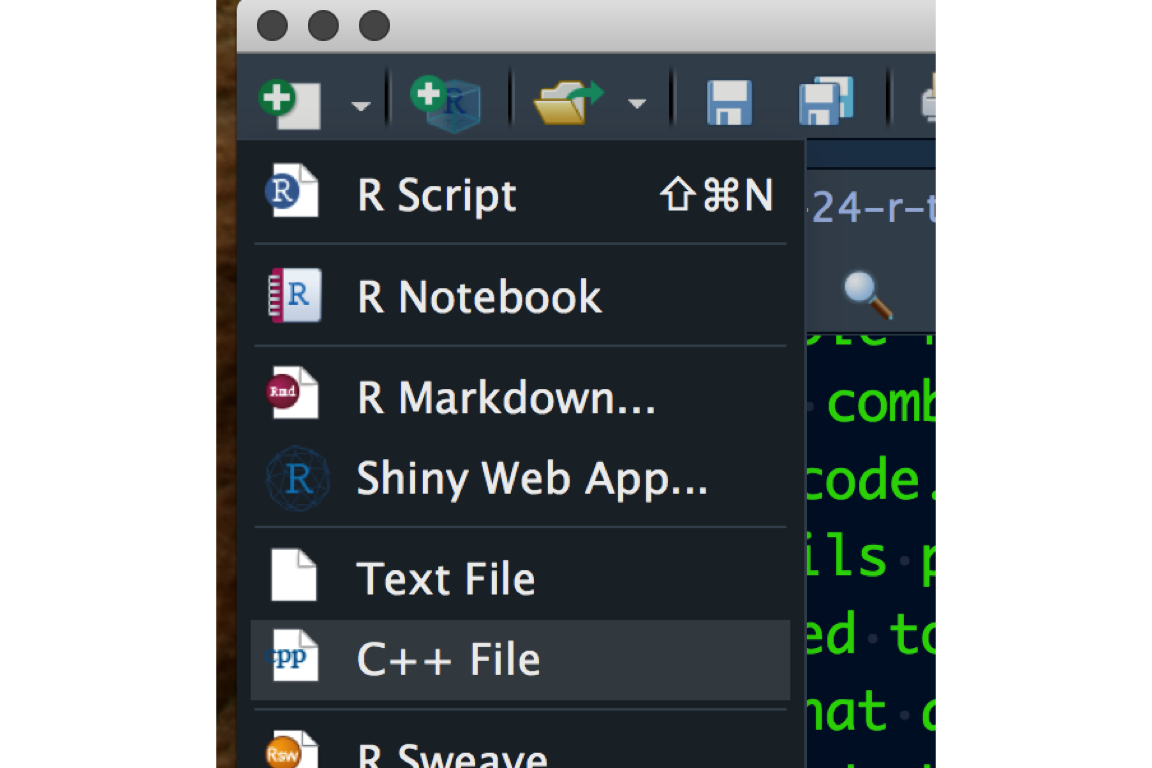
When you create a new RMarkdown document, you need to select the type of document you want to produce. Don’t worry, this option can be changed later:
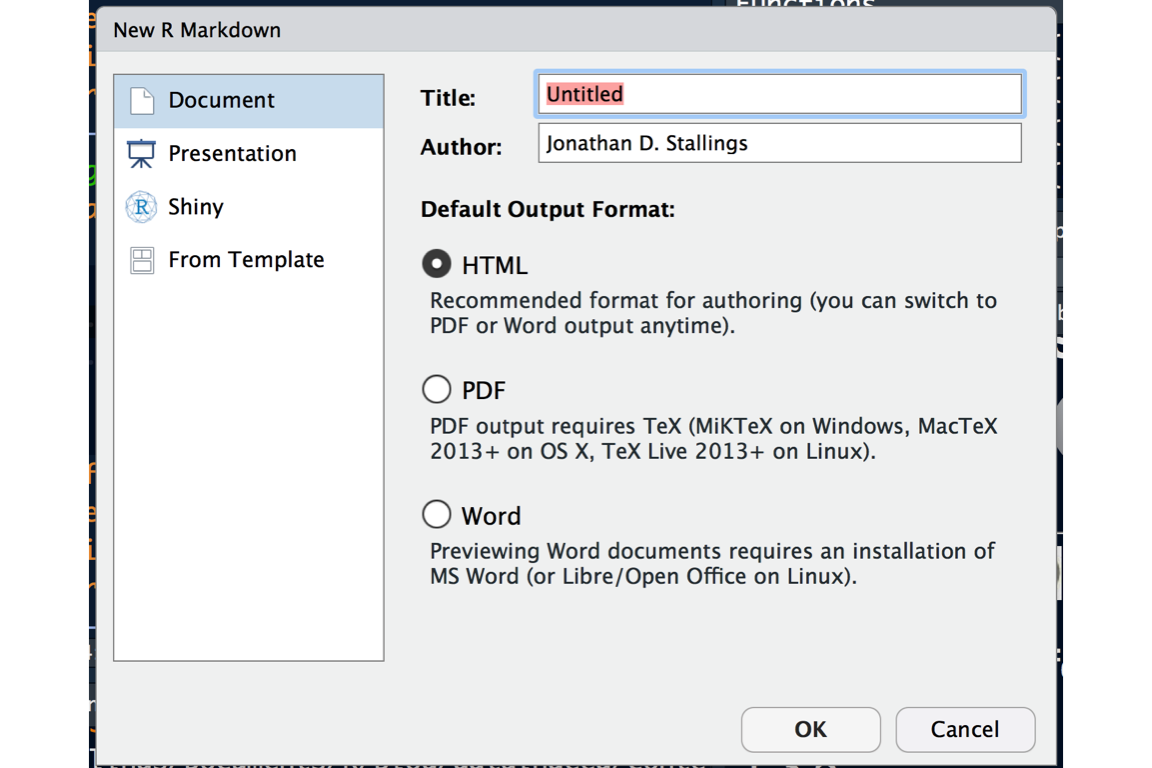
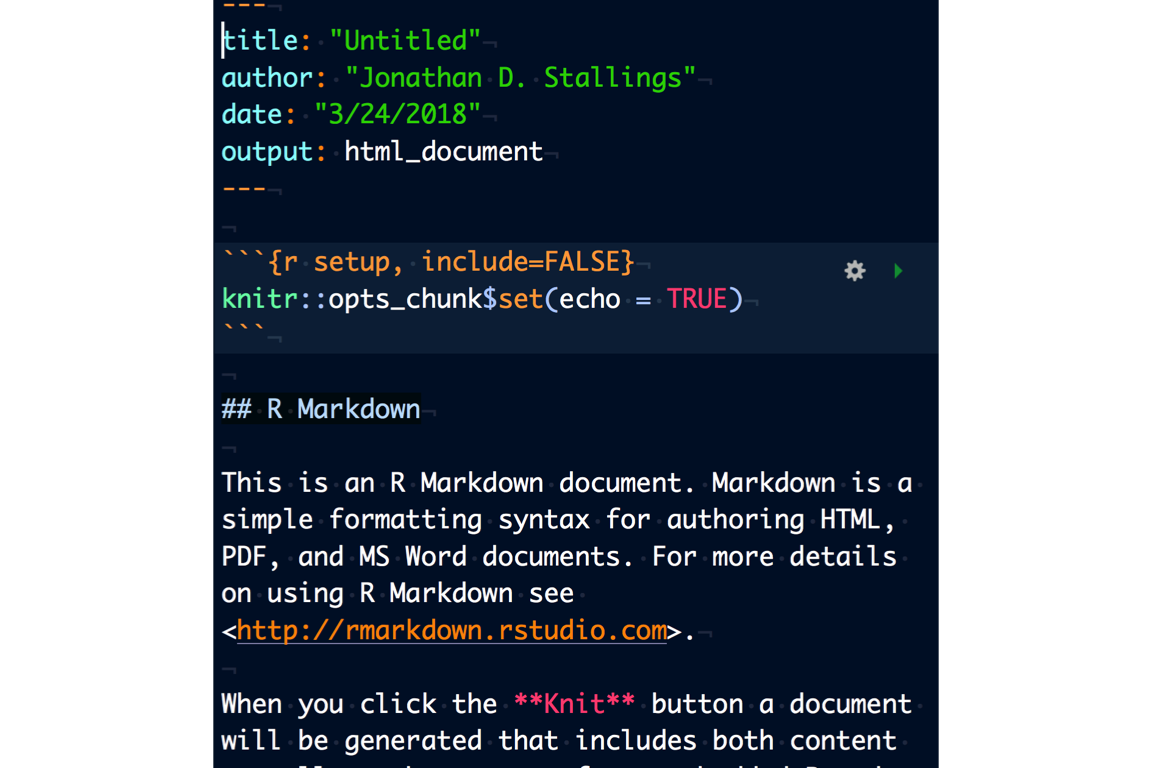
After you create the RMarkdown document, a basic template is created. The three main components of the document are:
YAML (separated by “—”)
Text (markdown, “#’s” represent various styles)
Code chunks (R, Python, SQL, etc…separated by “```”)
YAML Content
YAML contains all the metadata for your RMarkdown document. There are tons of different YAML document-specific features that can be reviewed here, but not all features are usable with the theme that you use to publish documents to a website. If your manuscript or data story is published to RStudio connect, no worries. Here are the YAML settings I use to make sure citations and CSL are incorporated in the knitted document.
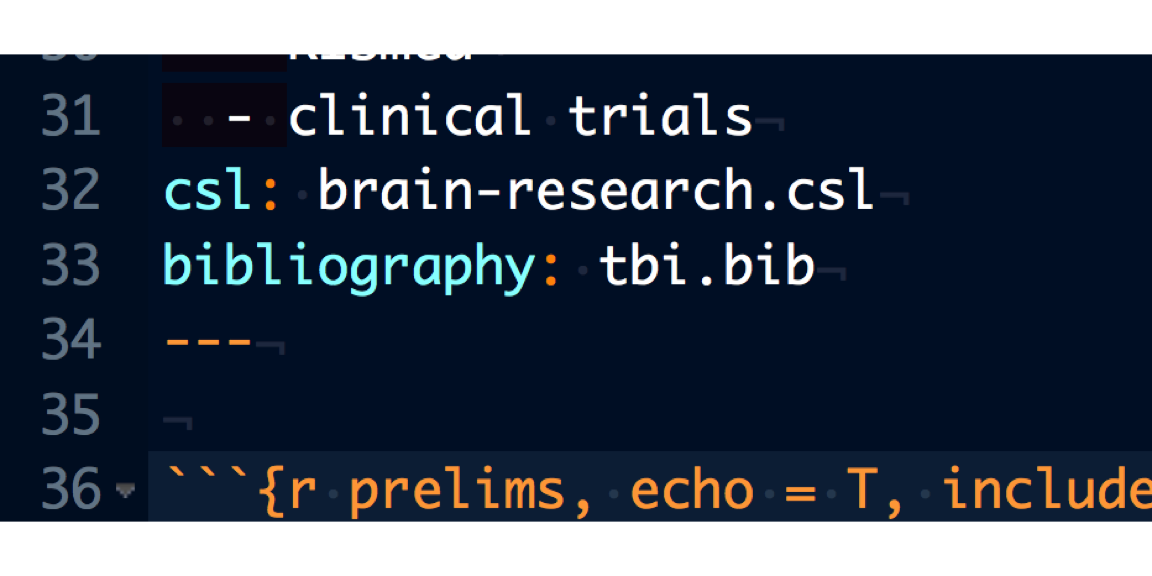
Citation Style Language
One example of information that is defined in the YAML is the citational style language (CSL). A great resource for CSL can be found here. Place the .csl file in the same folder location as the RMarkdown document.
Bibliography
The source of the bibliography is also defined in the YAML. I use both windows (for government work) and mac (all of my personal/business work), and unfortunately my approach to the bibliography is different on each system. On the mac, I recommend installing BibDesk and exporting your citation library as a .bib file. In the video below, I show how I import citations using BibDesk. You can download the citation from the site of interest, or search the sites of interest within BibDesk. Either way, when your library is complete, save it to the folder with your RMarkdown document. On windows, I use EndNote and export the library to a xml file.
{{ instagram Bgts-f5FgHe }}
Example Text with Citations
In-line Citations
More than 1.7 million TBIs occur each year in the United States , leading to 50,000 deaths.Faul and Coronado (2015) Many survivors suffer from TBI-related disabilities; approximately 5.3 million Americans require help in performing daily activities.Kabadi and Faden (2014) As a result, the healthcare burden of TBI in the U.S. exceeds $76 billion a year.Kabadi and Faden (2014)
Citations are embedded into the text with the @ symbol within brackets with the key names for each reference separated by commas:

Example Code Chunk
To insert a code chuck, select the insert button, which allows you to select a code chunk into the RMarkdown document. In the picture below, I show you the drop down screen and the code chunk that I use to call the image. Alternatively, you could use a custom function to incorporate a custom code chuck (see bonus material at the end of this document for details):
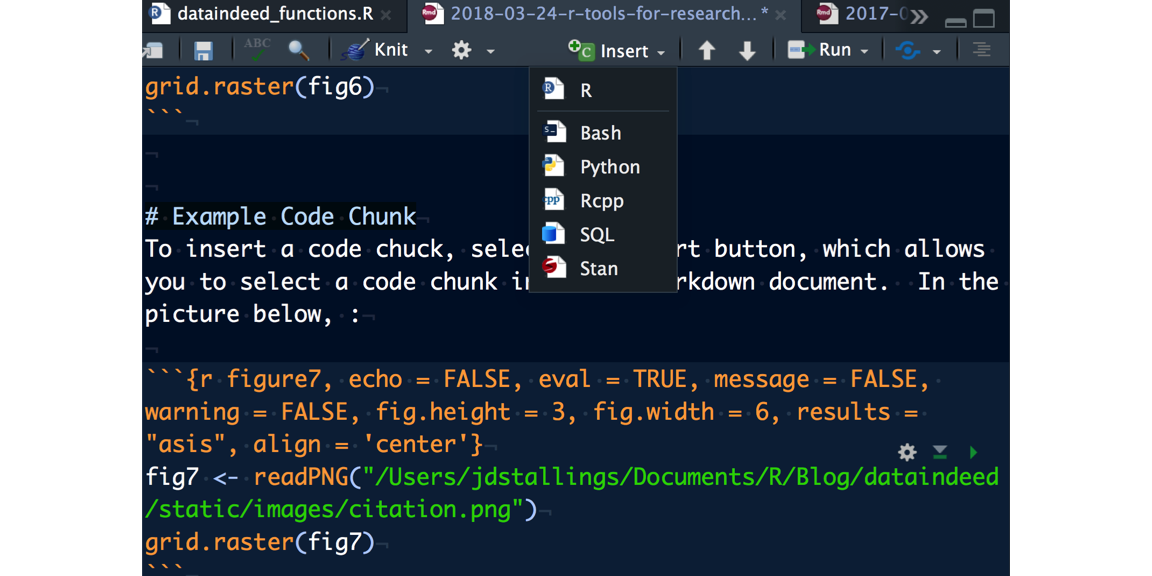
References
Here is the code chunk I use to incorporate the references in this document. Note that the actual references show up at the end of the document!
bibliography()Bonus Info:
Mathjax
If you post your manuscript or story to a website and require formulas to display, you need to incorporate mathjax into the RMarkdown document. Here is a basic tutorial on Mathjax. For my set up, I place the following code snippet in the footer.html layout for my website theme:
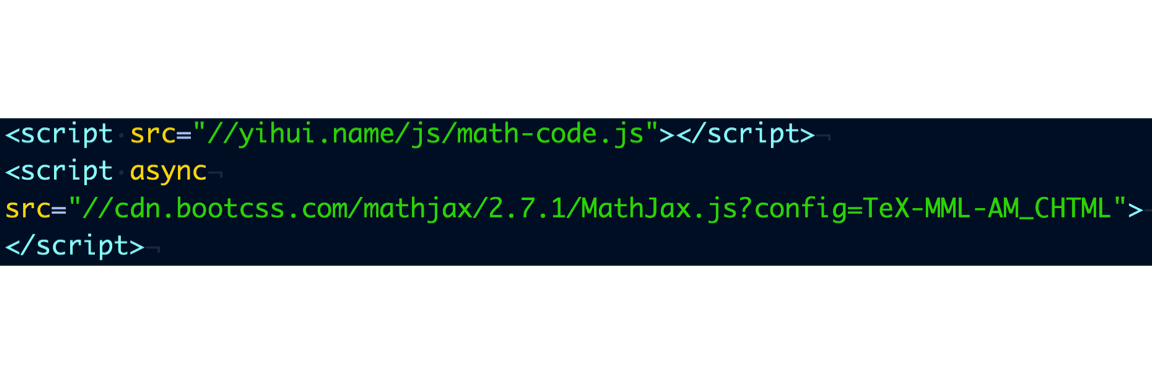
When you author your RMarkdown documents, ensure that you escape your equations with dollar signs, such as \(\sigma = \sum_{n-1}^i\) at the beginning and end of the inline equation.
For block equations, use double dollar signs: \[\sigma = \sum_{n-1}^i\]
Here is a photo of what it looks like written in the RMarkdown document:

Thanks for checking out my blog! Until next time…
